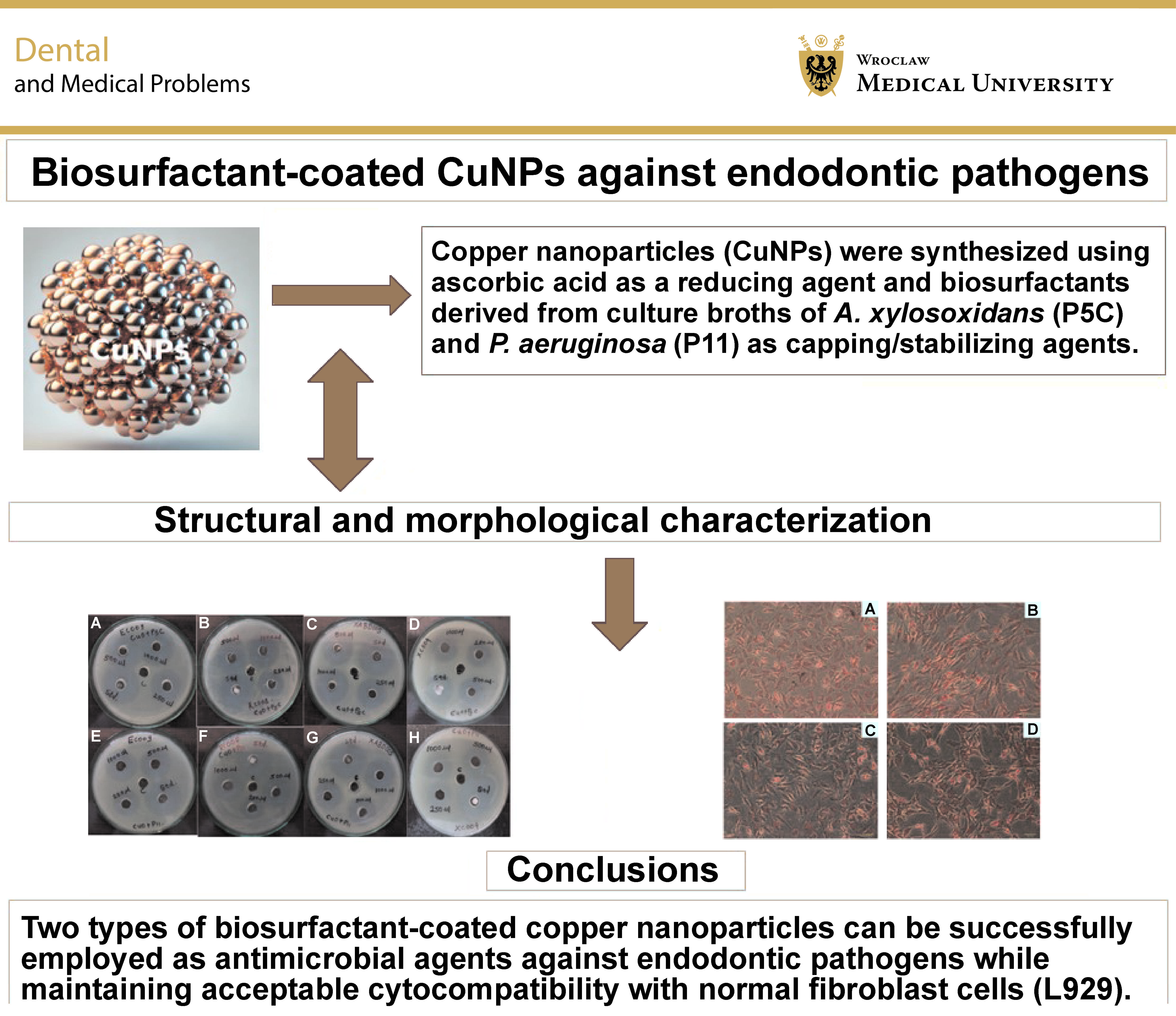
Abstract
Background. Endodontic infections caused by pathogens such as Escherichia coli, Staphylococcus aureus, Streptococcus oralis, and Enterococcus spp. pose significant challenges due to biofilm formation and resistance to conventional treatments. Copper nanoparticles (CuNPs) exhibit promising antimicrobial properties, and coating them with biosurfactants may enhance stability while reducing cytotoxicity.
Objectives. The aim of the study was to synthesize and characterize CuNPs stabilized with biosurfactants derived from Achromobacter xylosoxidans and Pseudomonas aeruginosa, and to evaluate their antimicrobial efficacy and cytotoxicity.
Material and methods. Copper nanoparticles were synthesized using a chemical reduction method and coated with biosurfactants to prevent oxidation and aggregation. The nanoparticles were characterized using Fourier transform infrared spectroscopy (FTIR), ultraviolet–visible (UV–Vis) spectroscopy, scanning electron microscopy (SEM), transmission electron microscopy (TEM), and energy-dispersive X-ray spectroscopy (EDX). Antimicrobial activity against E. coli, S. aureus, S. oralis, and Enterococcus spp. was evaluated using the well diffusion method and the minimum inhibitory concentration (MIC). Cytotoxicity was assessed using an MTT assay on L929 fibroblast cell lines.
Results. Characterization confirmed the formation of spherical CuNPs with sizes ranging from 90 nm to 100 nm. Both CuO+P5C and CuO+P11 nanoparticles demonstrated potent antimicrobial activity against all tested pathogens, with a MIC value of 125 µg/mL. Cytotoxicity results showed minimal toxicity, with nearly 78–80% cell viability maintained at concentrations of up to 100 µg/mL, indicating excellent biocompatibility.
Conclusions. Biosurfactant-coated CuNPs represent a stable, eco-friendly and effective antimicrobial alternative for addressing persistent endodontic infections. Further in vivo studies and extended cytotoxicity evaluations are needed to confirm their clinical application.
Keywords: copper nanoparticles, biosurfactants, Achromobacter xylosoxidans, Pseudomonas aeruginosa, endodontic pathogens, antimicrobial activity
Introduction
Endodontic infections are frequently caused by microbial species such as Escherichia coli, Staphylococcus aureus, Streptococcus oralis, and Enterococcus spp., which produce virulence factors like biofilms and toxins, complicating the outcomes of root canal treatment.1, 2, 3 These pathogens thrive in the nutrient-rich, anaerobic environment of infected root canals, contributing to tissue destruction and the formation of periapical lesions.4, 5 A comprehensive understanding of the resilience and virulence of these microorganisms is critical for the development of targeted strategies aimed at promoting healing, preventing reinfection and improving patient outcomes.6
Nanotechnology offers an innovative platform to combat resistant microorganisms by employing nanoscale materials (1–100 nm) with high surface areas and enhanced reactivity.7 Among various nanoparticles, copper nanoparticles (CuNPs) are particularly notable for their ability to release reactive ions that disrupt microbial membranes.8, 9 These properties position CuNPs as promising alternatives to conventional antibiotics, particularly in challenging biomedical applications such as endodontic therapy. Their small size and high surface-to-volume ratio facilitate effective penetration into biofilms and microbial membranes, making them highly effective antimicrobial agents.10, 11
Historically, copper compounds have been widely used in agriculture for their bactericidal properties. However, recent advancements have focused on the application of CuNPs in the biomedical field. Coating CuNPs with biosurfactants enhances their stability and reduces cytotoxicity, broadening their potential applications.12 Biosurfactants are amphiphilic compounds produced by microorganisms such as bacteria, fungi and yeasts, and are known for their eco-friendly, biodegradable nature and surface-active properties. These compounds reduce interfacial tension, act as natural capping agents that stabilize nanoparticles, and prevent aggregation and oxidation.13, 14, 15 The functional groups present in biosurfactants enable controlled nanoparticle formation and facilitate effective interactions with microbial cell walls, thereby enhancing antimicrobial efficacy.16, 17
Biosurfactants, including glycolipids such as rhamnolipids derived from Achromobacter xylosoxidans and Pseudomonas aeruginosa, exhibit excellent emulsifying properties. These characteristics allow them to form coatings on CuNPs, enhancing dispersibility, biocompatibility and antimicrobial activity while reducing cytotoxicity.14 Given the microbial challenges associated with root canal infections, biosurfactant-coated CuNPs represent a novel, eco-friendly approach to improving treatment outcomes.
Objectives
The objective of this study was to compare the antimicrobial activity of CuNPs coated with biosurfactants derived from gram-negative endodontic bacteria. By leveraging recent advancements in biosurfactant-coated nanoparticles, this research aimed to explore their potential as effective alternatives to traditional antimicrobial agents for managing endodontic infections.
Hypotheses
The following hypothesis was formulated: Biosurfactant-coated CuNPs exhibit enhanced antimicrobial activity and reduced cytotoxicity compared to uncoated CuNPs when tested against endodontic pathogens. The null hypothesis states that biosurfactant-coated CuNPs do not significantly differ from uncoated CuNPs in terms of antimicrobial efficacy or cytotoxicity.
Material and methods
Chemicals and reagents
Copper sulfate pentahydrate and ascorbic acid were purchased from Sigma-Aldrich (St. Louis, USA). Deionized water was used to prepare the copper sulfate solution and plant extract. Sodium hydroxide, nitric acid and sodium bicarbonate, used in the synthesis process, were procured from Qualigens Pharma Pvt. Ltd. (Khalapur, India).
Synthesis of CuNPs
Copper nanoparticles were synthesized using a chemical reduction method as described by Ahamed et al.18 and Chowdhury et al.19 A 0.1 M solution of copper sulfate pentahydrate was prepared and stirred for 30 min. Then, 50 mL of 0.1 M ascorbic acid was added as a reducing agent, followed by 30 mL of 1 M sodium hydroxide for pH adjustment. The reaction was carried out at 80°C for 2 h, during which the color of the solution changed from yellow to reddish-brown, indicating the formation of CuNPs. The resulting product was left to settle for 24 h, washed with deionized water and ethanol, and stored at 4°C for further analysis.
Production of biosurfactants from microorganisms
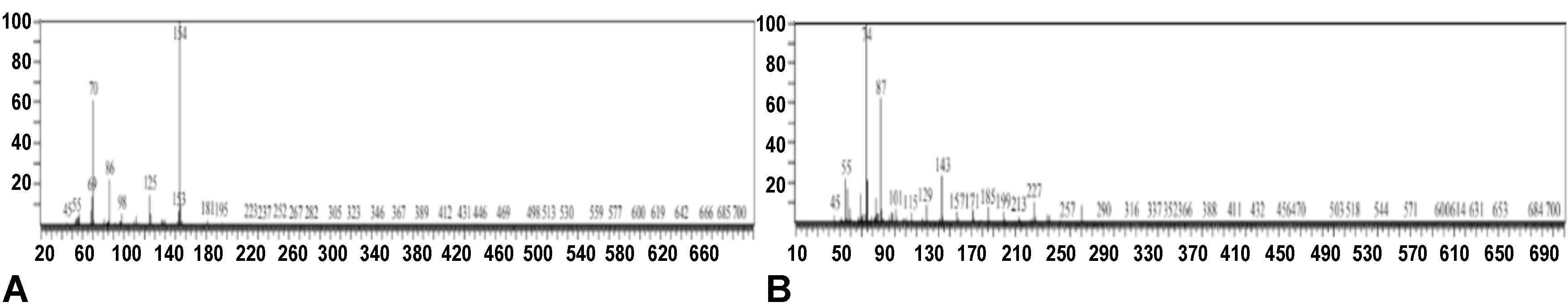
Mineral salt medium (MSM) containing 2% glucose and adjusted to pH 7 was prepared and sterilized in a 250-mL Erlenmeyer flask. Subsequently, 5 mL of culture broth from A. xylosoxidans (P5C) or P. aeruginosa (P11), each at a cell concentration of 1 × 109 cells/mL, was inoculated into the medium, along with 2 mL of petrol. The flask was incubated on an orbital shaker at 120 rpm and 35°C for 3 days. The culture broth was then centrifuged at 15,000 rpm and 4°C for 10 min to obtain a cell-free supernatant, which was filtered through a 0.2-µm Whatman filter paper. The supernatant was acidified to pH 2 using 6 N HCl, extracted with an organic solvent, and evaporated at 45°C (Table 1). The remaining solutes were dissolved in a solvent, and the biosurfactant stock was maintained at 1 mg/mL. Fourier transform infrared spectroscopy (FTIR) and liquid chromatography–mass spectrometry (LC–MS) analyses confirmed the biosurfactant composition.20
Coating of copper nanoparticles with biosurfactants
The pre-synthesized CuNPs were dissolved in 10 mL of deionized water and stirred with 200 ppm biosurfactant solution. Then, 2 M nitric acid was added dropwise to adjust the pH. A 1:1 ratio of the CuNP solution to biosurfactant (P5C or P11) dispersed in 50 mL of sodium bicarbonate was prepared. The reaction mixture was incubated at 37°C for 48 h, centrifuged at 2,500 rpm for 15 min, and the resulting biosurfactant-coated CuNPs were washed with deionized water and ethanol before being stored at 4°C.21
Characterization of copper nanoparticles
Ultraviolet–visible spectroscopy analysis
The optical properties of the nanoparticles were analyzed using an ultraviolet–visible (UV–Vis) spectrophotometer (UV-1800; Shimadzu, Kyoto, Japan). The absorbance peaks were measured to confirm nanoparticle formation. The suspension was prepared in a suitable solvent, after which the absorbance was evaluated.22
Fourier transform infrared spectroscopy analysis
Fourier transform infrared spectroscopy analysis (Nicolet iS10; Thermo Fisher Scientific, Waltham, USA) was used to identify functional groups associated with the nanoparticles and the biosurfactant coating. The minimum required quantity of samples was loaded onto the sampler.22
Zeta potential analysis
Zeta potential was measured using a Zetasizer Nano ZS (Malvern Panalytical, Malvern, UK) to evaluate nanoparticle stability. Deionized water was used to prepare the sample suspension, and the sample was injected into the sample holder to prevent air bubbles. Subsequently, the sample holder was retained inside the apparatus, and the zeta potential of the particles was analyzed using the electrophoretic light scattering method.22
Scanning electron microscopy and transmission electron microscopy analyses
Morphological characteristics were examined using scanning electron microscopy (SEM) (JSM-6390LV; JEOL Ltd., Tokyo, Japan) and transmission electron microscopy (TEM) (Tecnai G2 Spirit BioTWIN; FEI, Hillsboro, USA). Copper nanoparticles were dissolved in an appropriate solvent to prepare the sample suspension, which was then sonicated for 1 min. A sample drop was placed on the conductive substrate. Once the material had dried, the samples were analyzed using a TEM instrument set to 80–200 keV and an SEM device set to 10–20 keV.22
Energy-dispersive X-ray spectroscopy analysis
The elemental composition was determined using energy-dispersive X-ray spectroscopy (EDX) by depositing CuNPs on a carbon film and analyzing them under a microscope (JSM-IT300; JEOL Ltd.).23
Antimicrobial activity
Gram-positive pathogens, including S. aureus, S. oralis and Enterococcus spp., as well as gram-negative pathogens, including E. coli, were procured from the CSIR–National Collection of Industrial Microorganisms (NCIM) (Pune, India) and inoculated onto nutrient agar plates at 37°C. The antimicrobial activity of CuNPs was determined against 4 different endodontic pathogens, namely E. coli (EC009), S. aureus (RC008), S. oralis (XaBDOO5), and Enterococcus spp. (XCOO4) using a 96-well plate method.24, 25, 26, 27 These gram-negative, rod-shaped bacteria cause pathogenic diseases in many plants. Depending on the host plant and specific strain, they can lead to leaf spots, cankers, wilting, stunting, and fruit blemishes. In the present study, the method used was the resazurin assay.28, 29
Well diffusion method
Nutrient agar (15 mL) was poured into sterilized Petri dishes, inoculated with 100 µL of bacterial broth, and spread evenly. Wells with a diameter of 6 mm were created in the agar using a sterile borer, and 100 µL of the CuNP solution (100 µg/mL) was added. The plates were incubated at 37°C for 24 h, and the diameter of the zone of inhibition (ZOI) was measured. Streptomycin (1 mg/mL) and dimethyl sulfoxide (DMSO) were used as positive and negative controls, respectively.
Minimum inhibitory concentration analysis
The minimum inhibitory concentration (MIC) was determined using the resazurin assay in a 96-well plate. Serial dilutions of CuNPs were prepared, and 100 µL of each concentration was added to the wells, followed by 10 µL of bacterial suspension and 50 µL of nutrient broth. After incubation at 37°C for 18–24 h, MIC values were assessed by observing the color change of resazurin from blue to pink.24, 25 The percentage of inhibition was calculated using the following formula (Equation 1):
Cytotoxicity analysis over normal human fibroblast cell lines
Normal fibroblast cells (L929) were procured from the National Centre for Cell Science (NCCS), Pune, India. The cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM) (Gibco™, Thermo Fisher Scientific), supplemented with 10% fetal bovine serum (FBS) (Sigma-Aldrich) under standard conditions (37°C, 5% CO2). The cells were seeded at a density of 1 × 104 cells/well in a 96-well plate and incubated for 24 h. They were then treated with CuNPs (20, 40, 60, 80, 100 µg/mL) for 24 h. Subsequently, 10 µL of the MTT solution (5 mg/mL in phosphate-buffered saline (PBS)) was added and incubated for 3–4 h at 37°C. After removing the medium, 100 µL of DMSO was added to dissolve the formazan crystals, and the absorbance was measured at 570 nm.30 Cell viability was calculated as follows (Equation 2):
where:
OD – optical density.
Statistical analysis
All experiments were performed in triplicate, and the results were expressed as mean ± standard deviation (M ±SD). The statistical analysis was conducted using GraphPad Prism 9.0 software (GraphPad Software, Inc., San Diego, USA). Differences between experimental groups were evaluated using one-way analysis of variance (ANOVA), followed by Tukey’s post hoc test for pairwise comparisons. A p-value <0.05 was considered statistically significant.
Results
Characterization of biosurfactants
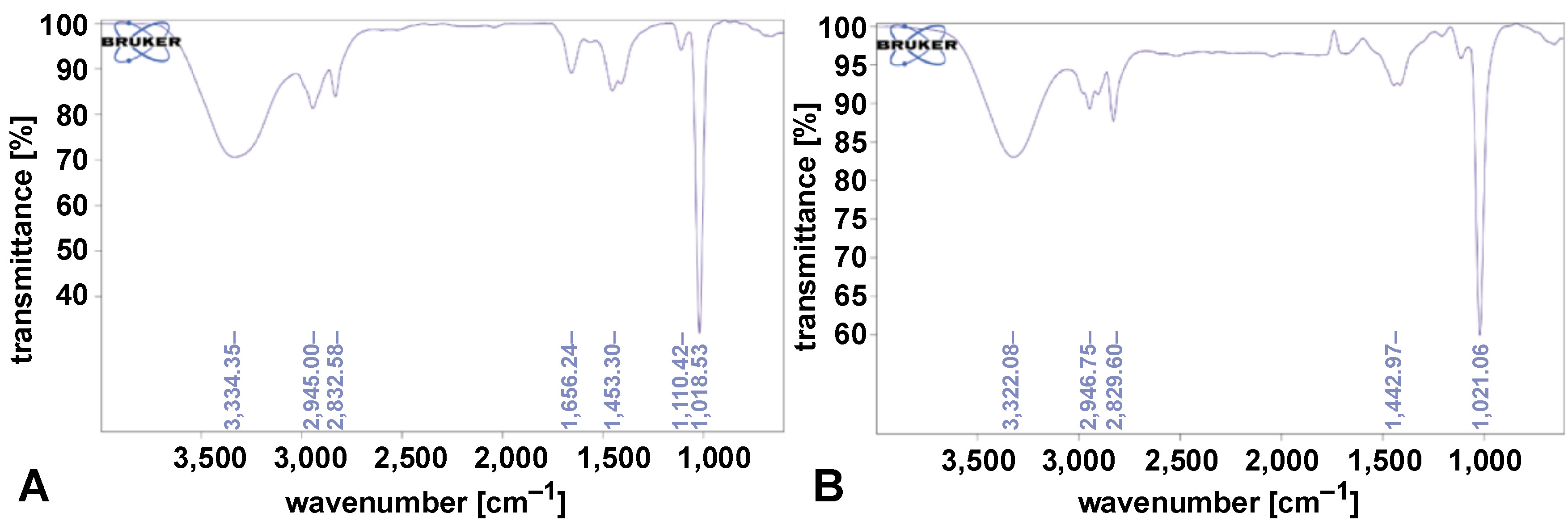
The FTIR analysis of the biosurfactants showed characteristic peaks confirming their composition. Peaks corresponding to proteins were observed at 1,656.24 cm−1, carbohydrates at 1,110.42 cm−1 and 1,018.53 cm−1, and fatty acids at 2,832.58 cm−1 and 2,945.00 cm−1. A broad O–H stretching peak was identified at 3,334.35 cm−1, indicating hydroxyl groups. These spectra confirmed the biosurfactants produced by A. xylosoxidans and P. aeruginosa, as shown in Figure 1 and Figure 2.
Characterization of CuNPs
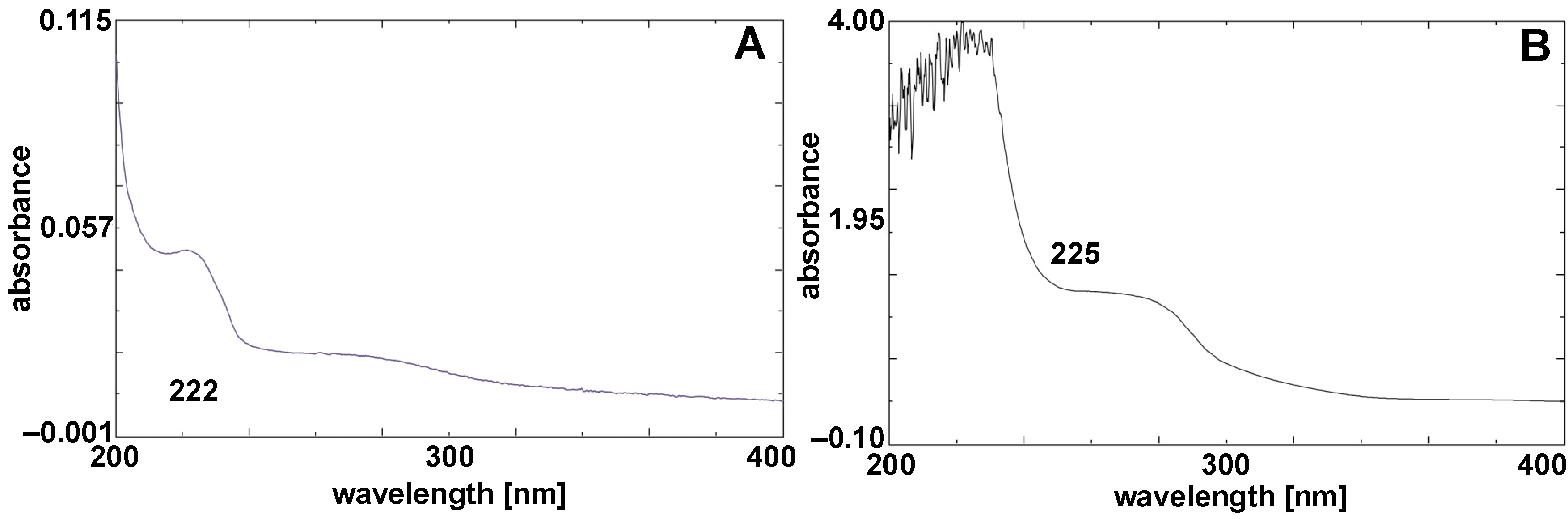
(UV–Vis spectroscopy)
UV–Vis spectroscopy confirmed the formation of CuNPs, with maximum absorbance peaks observed at 222 cm−1 and 225 cm−1. These peaks are characteristic of biosynthesized CuNPs, as presented in Figure 3.
FTIR analysis for CuNPs
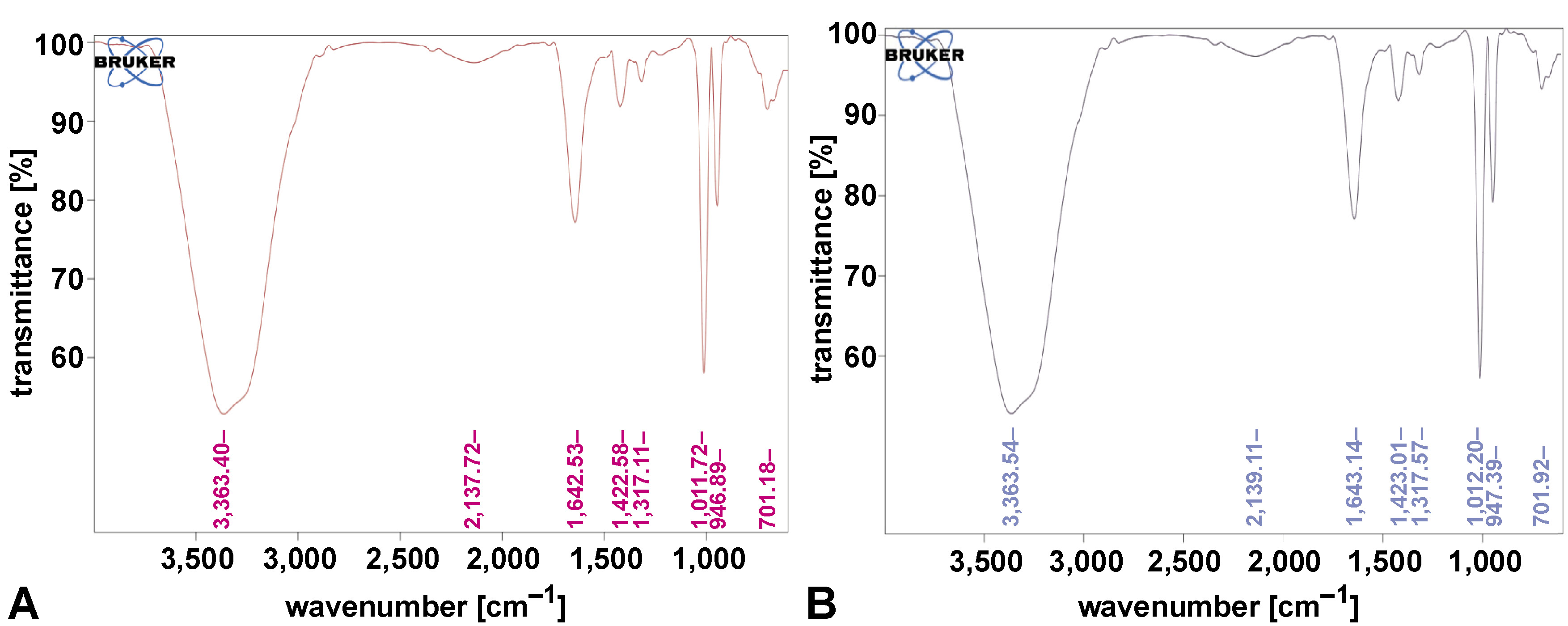
The FTIR spectra of CuNP samples revealed key functional groups associated with the nanoparticles. Peaks at 701 cm−1 corresponded to Cu–O stretching, confirming the presence of copper oxide. O–H stretching was observed at 3,363.40 cm−1, and C≡C stretching was identified at 2,137.72 cm−1. A peak at 1,643.14 cm−1 indicated C=O stretching vibrations, while 1,422.58 cm−1 corresponded to C–H bending. These findings confirm the successful synthesis and capping of CuNPs, as shown in Figure 4.
Zeta potential analysis
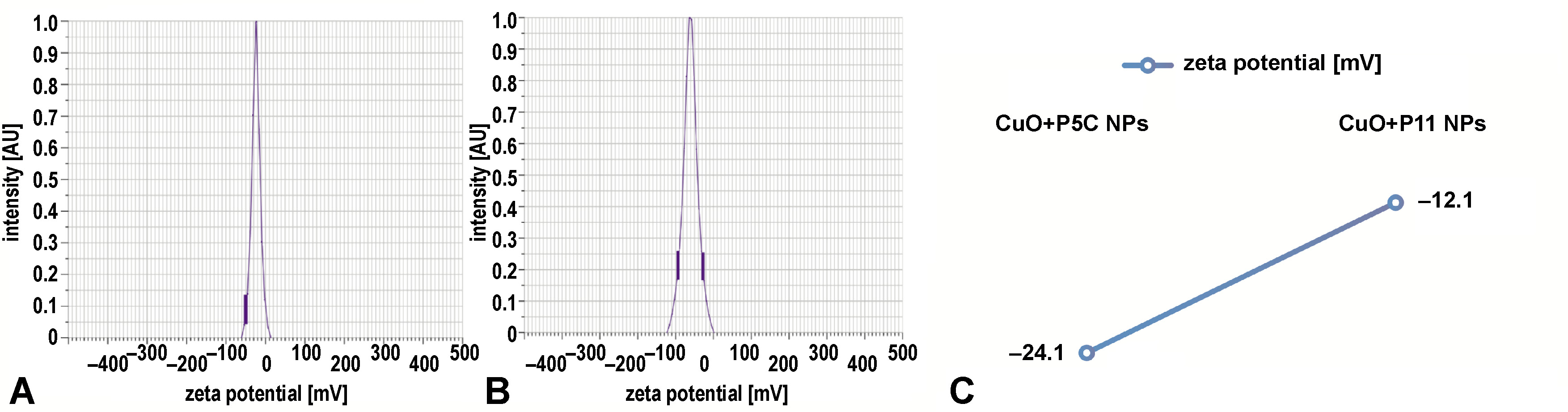
Zeta potential measurements were conducted to evaluate the stability of CuNPs. The CuO+P5C nanoparticles exhibited a zeta potential of −24.1 mV, while the CuO+P11 nanoparticles showed a zeta potential of −12.1 mV, indicating moderate stability. These moderately negative charges suggest reduced toxicity and enhanced biological compatibility. Figure 5 displays zeta potential distributions for both nanoparticle types.
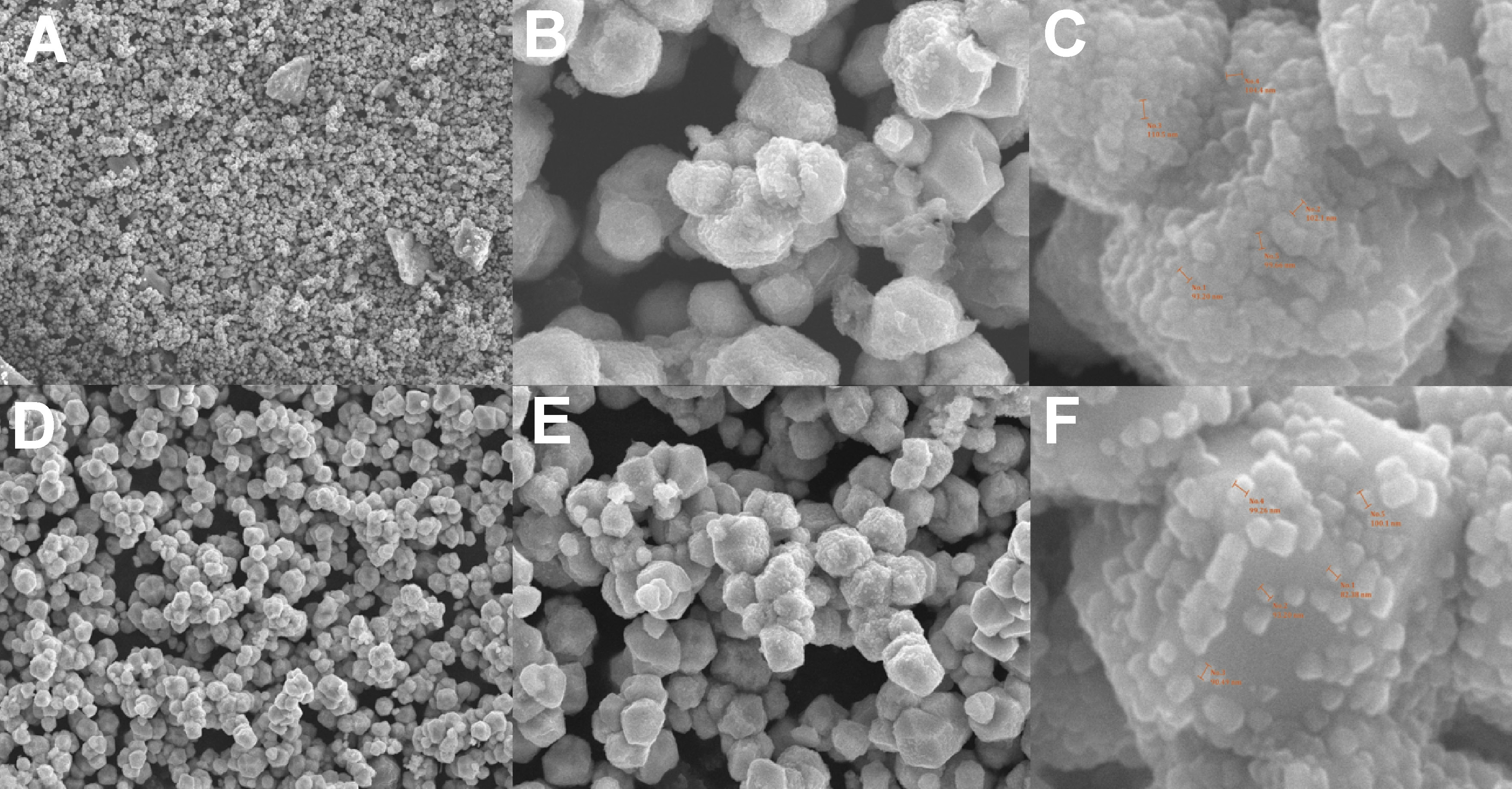
SEM analysis
Scanning electron microscopy revealed distinct morphological characteristics of CuNPs. The CuO+P5C nanoparticles exhibited irregular morphology with sizes ranging from 90 nm to 100 nm, whereas the CuO+P11 nanoparticles displayed more granular and spherical morphology with sizes between 85 nm and 95 nm. Full-field SEM images are presented in Figure 6.
Morphological analysis by TEM
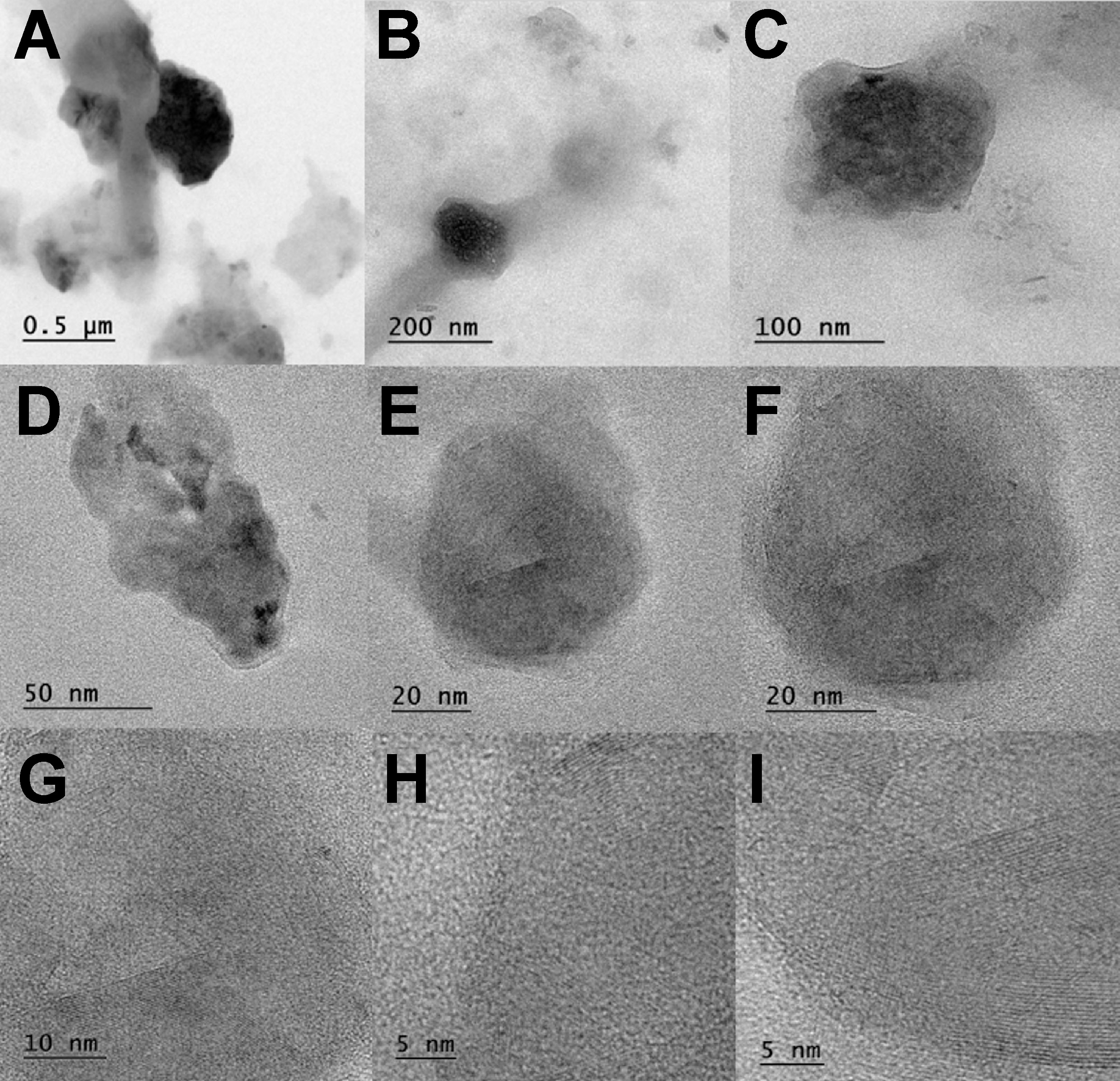
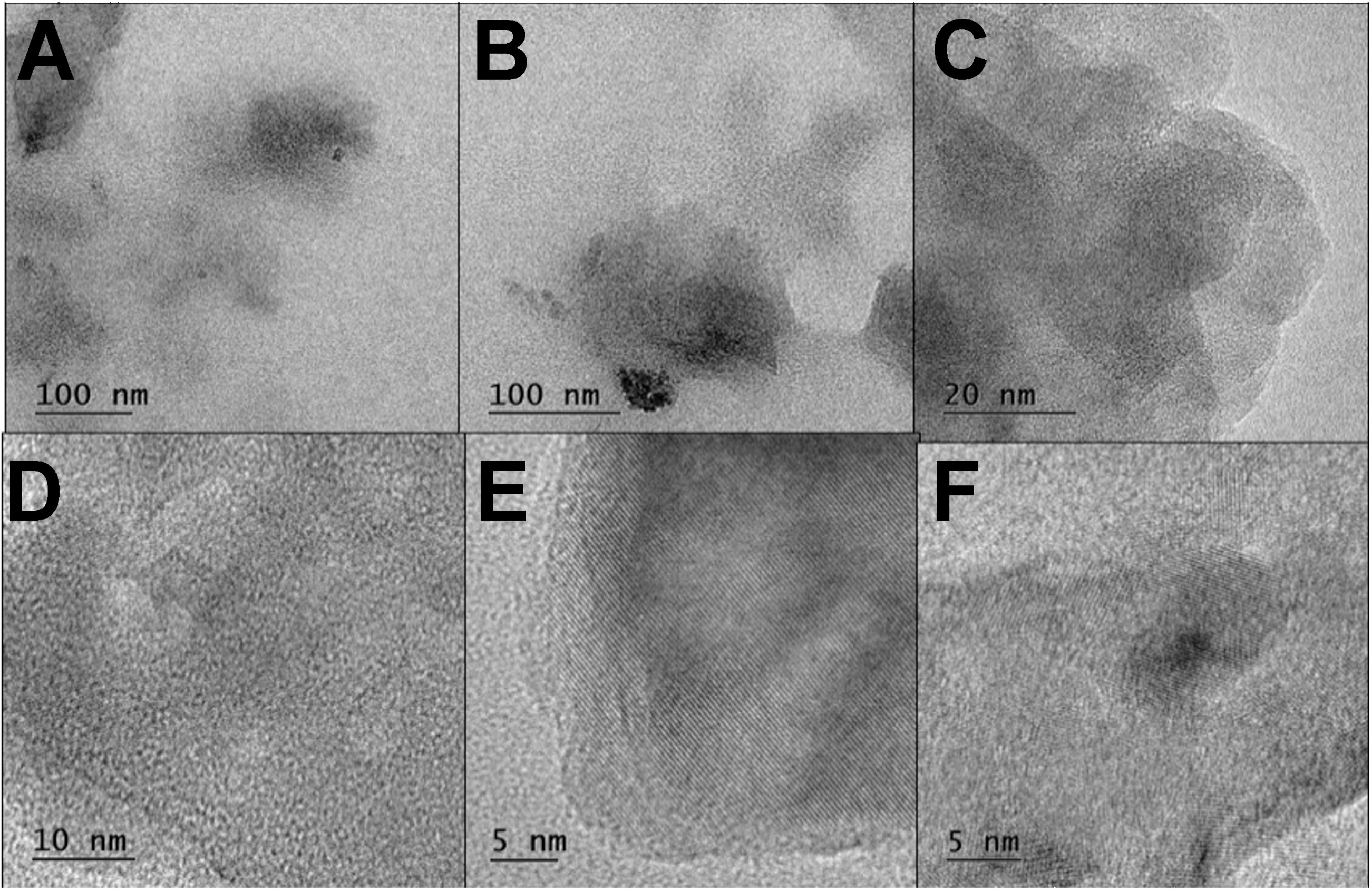
Transmission electron microscopy images confirmed the spherical shape and size distribution of CuNPs. The nanoparticles ranged in diameter from 90 nm to 100 nm. Figure 7 and Figure 8 illustrate the spherical morphology and size distribution of CuNP samples.
Composition analysis by EDX
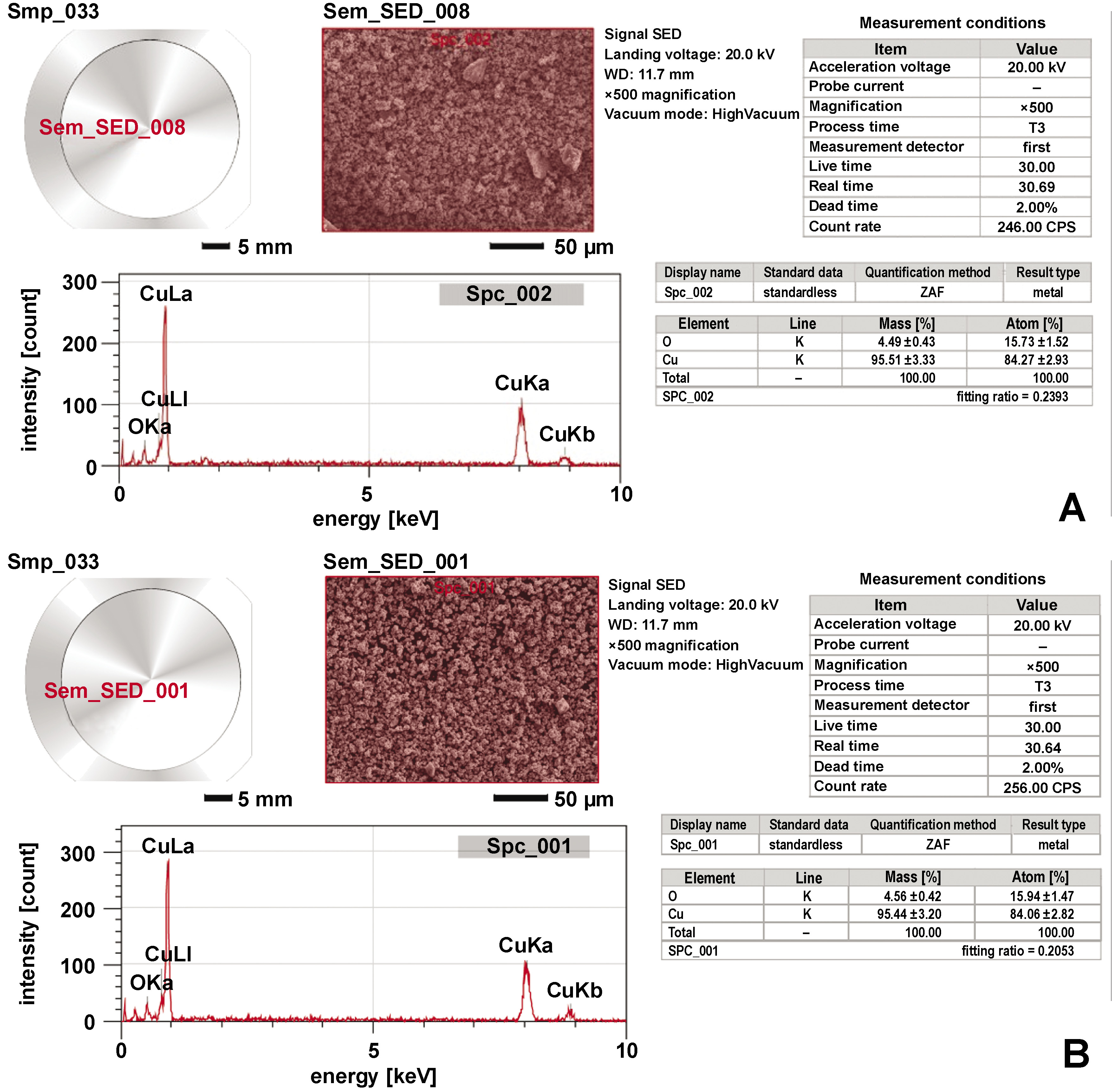
Energy-dispersive X-ray spectroscopy confirmed the elemental composition of the nanoparticles. Copper accounted for 95.51 ±3.33% of the total weight in the CuO+P5C samples and 95.44 ±3.20% in the CuO+P11 samples. Prominent peaks at 8.04 keV further validated the high copper content in the nanoparticles (Figure 9).
Antimicrobial activity of CuNPs
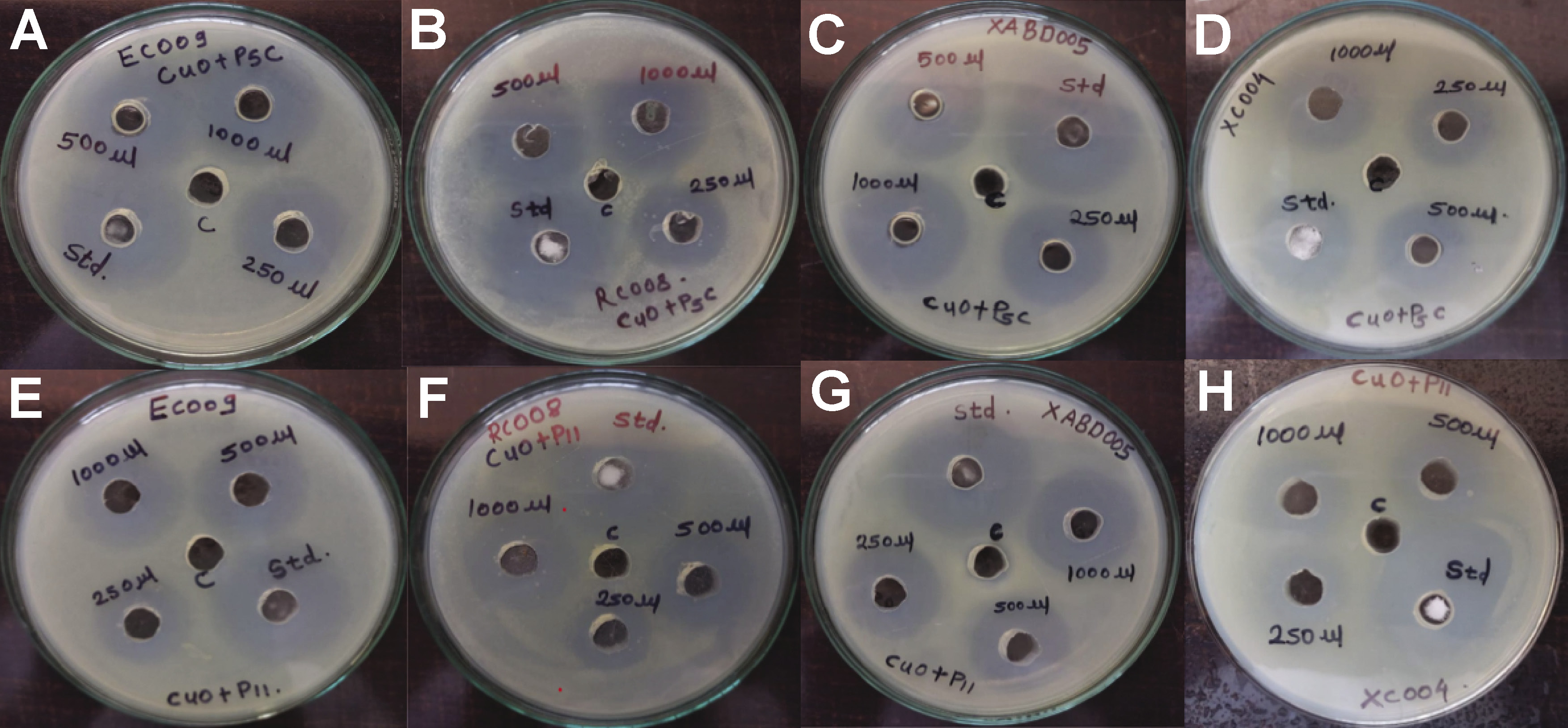
The antimicrobial efficacy of CuNPs was assessed against E. coli, S. aureus, S. oralis, and Enterococcus spp. The ZOI results, summarized in Table 2 and Figure 10, indicated significant activity in both the CuO+P5C and CuO+P11 samples. Escherichia coli exhibited the highest sensitivity to the nanoparticles. Streptomycin was used as a positive control. Biosurfactant-coated CuNPs demonstrated potent antimicrobial activity against all tested strains.
Minimum inhibitory concentration analysis
The MIC of both CuO+P5C and CuO+P11 nanoparticles was found to be 125 µg/mL for all tested microorganisms, as measured by the resazurin assay. The results, summarized in Table 3, confirm the strong antimicrobial potential of biosurfactant-coated CuNPs.
Cytotoxicity analysis
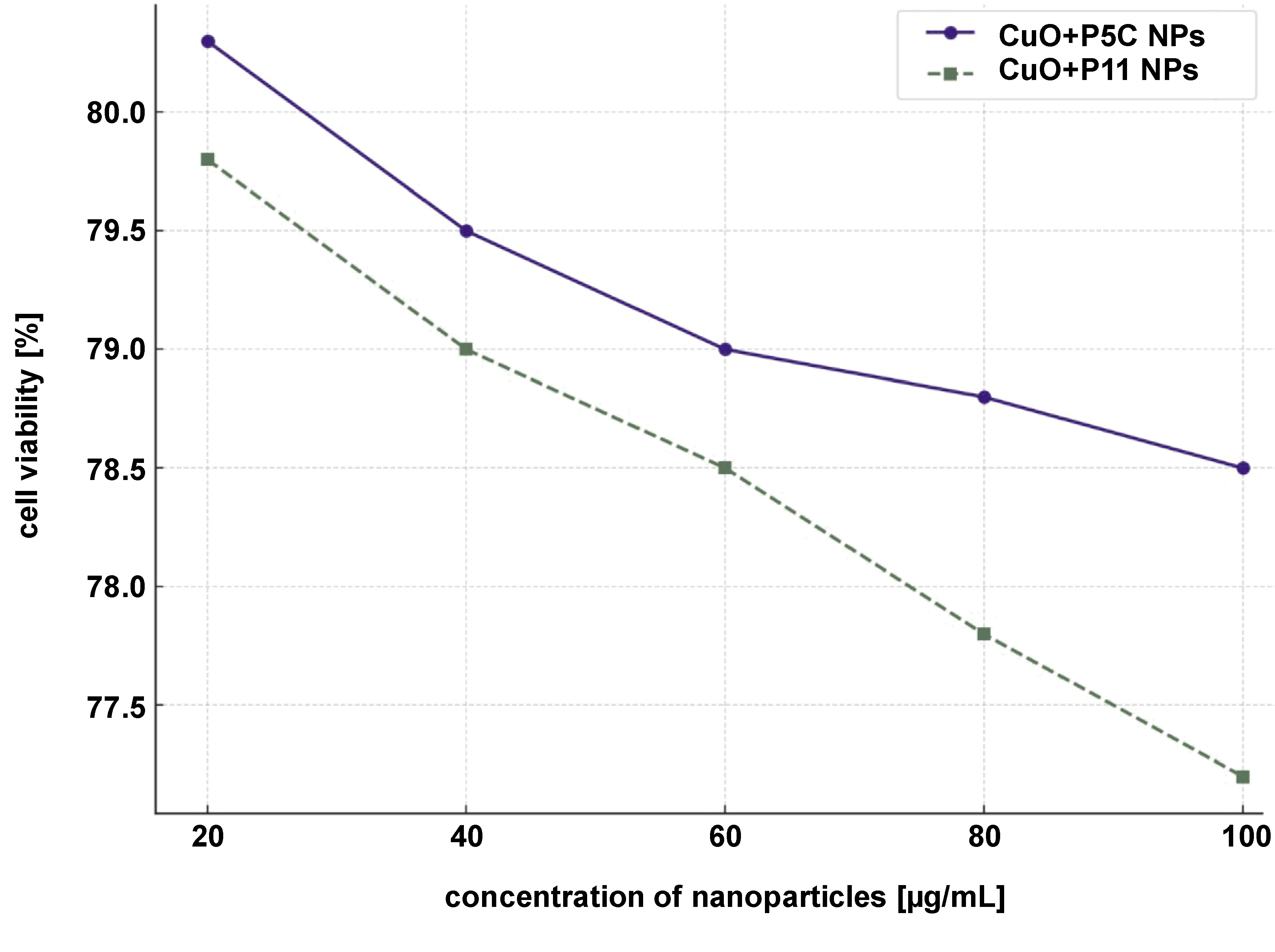
The cytotoxicity of CuNPs was assessed using the MTT assay on L929 fibroblast cell lines. The CuO+P5C and CuO+P11 nanoparticles exhibited good cytoprotective activity, maintaining nearly 78–80% of cell viability at concentrations up to 100 µg/mL. These results, compared to 70% ethanol as a positive control, indicate the superior biocompatibility of the nanoparticles (Figure 11).
Discussion
The present study demonstrates the successful synthesis and characterization of CuNPs stabilized with A. xylosoxidans (P5C) and P. aeruginosa (P11) biosurfactants. Fourier transform infrared spectroscopy analysis of nanoparticles provided insights into the functional groups on their surface, which are related to the coating agents used during synthesis. In the present study, the FTIR spectrum of biosurfactant-coated CuNPs revealed key functional groups associated with the biosurfactants. A broad peak at 3,363.40 cm−1 corresponds to O–H stretching, indicating hydroxyl groups involved in stabilization. The peak at 1,643.14 cm−1 is attributed to C=O stretching vibrations, likely from carbonyl groups in the biosurfactants. A peak at 701 cm−1 confirms Cu–O stretching, indicating the presence of copper oxide bonds. Additional peaks at 1,422.58 cm−1 and 2,137.72 cm−1 correspond to C–H bending vibrations and C≡C stretching, respectively. These findings confirm the successful coating of the nanoparticles with biosurfactants, which play a critical role in enhancing stability and preventing aggregation.
The SEM and TEM structural analyses revealed that the biosurfactant-coated CuNPs were spherical and granular, with particle sizes ranging from 90 nm to 100 nm. While both types of nanoparticles exhibited similar morphology, the granular structure of the CuO+P11 samples suggests subtle differences in stabilization properties. The zeta potential analysis revealed values of −24.1 mV (CuO+P5C) and −12.1 mV (CuO+P11), suggesting moderate stability and a slight variation in surface charge, which may influence their interaction with bacterial membranes and their cytotoxicity profiles. The size range (90–100 nm) and spherical morphology of the synthesized CuNPs align with previous studies reporting nanoparticle dimensions influenced by synthesis conditions and capping agents.20, 31 However, this study uniquely demonstrates the role of biosurfactant coating in enhancing nanoparticle stability and biocompatibility while retaining antimicrobial potency. The variations in size can be attributed to the differences in synthesis conditions, reaction medium compositions and experimental protocols. The size of CuNPs can span from a few nanometers to several hundred nanometers, illustrating the flexibility in manipulating their dimensions for diverse uses in catalysis, biomedicine and bioremediation.32
The UV–Vis spectrum of nanoparticles provides critical information about their surface plasmon resonance (SPR), a phenomenon arising from the collective oscillation of conduction electrons when nanoparticles interact with light at a specific wavelength. The observed absorbance peak indicates the successful formation of nanoparticles and is influenced by factors such as particle size, shape, coating, and surface modifications. In this study, the characteristic SPR peaks of biosurfactant-coated CuNPs at 222 nm and 225 nm confirm their formation. Furthermore, UV–Vis spectroscopy is a valuable tool for studying the interaction of nanoparticles with target molecules, as changes in SPR can reveal insights into nanoparticle stability, aggregation and surface interactions.33
The antimicrobial activity of CuNPs was evaluated against E. coli, S. aureus, S. oralis, and Enterococcus spp., which are commonly implicated in persistent endodontic infections. The CuO+P5C and CuO+P11 nanoparticles demonstrated significant activity, with a MIC value of 125 µg/mL for all bacterial strains. These results confirm that CuNPs effectively inhibit bacterial growth through metal ion release and membrane penetration mechanisms.34, 35 Notably, the highest sensitivity was observed in E. coli, likely due to its thinner cell wall structure compared to gram-positive bacteria.
The cytotoxicity analysis on L929 fibroblast cell lines revealed that CuO+P5C and CuO+P11 exhibited minimal toxicity at 100 µg/mL concentrations, with cell viability ranging from nearly 78% to 80%. This result indicates that biosurfactant coating significantly reduces nanoparticle toxicity while improving biocompatibility. These findings align with studies reporting that biosurfactants provide a natural, eco-friendly capping layer that reduces nanoparticle aggregation and oxidation while enhancing stability and biological safety.36, 37
The enhanced antimicrobial efficacy of biosurfactant-coated CuNPs can be attributed to several factors. Firstly, biosurfactant-mediated stabilization plays a critical role in preventing nanoparticle oxidation and aggregation, which enhances their stability and promotes effective interaction with microbial cell walls.38 Secondly, the small size and high surface area of nanoparticles facilitate better penetration through bacterial membranes and enable the generation of reactive copper ions, leading to cellular damage and microbial inhibition.39 Lastly, the moderately negative zeta potential of the nanoparticles improves their adhesion to bacterial membranes while minimizing excessive cytotoxic effects, thus ensuring a balance between antimicrobial activity and biocompatibility.40
The novelty of this study lies in the use of biosurfactants from A. xylosoxidans and P. aeruginosa, which have not been widely explored for nanoparticle synthesis. These findings open avenues for eco-friendly and cost-effective approaches to nanoparticle-mediated antimicrobial therapies, particularly for endodontic infections, where biofilm formation and microbial persistence pose significant treatment challenges.
Limitations and future directions
While the study demonstrated the potential of biosurfactant-coated CuNPs, several limitations should be acknowledged. The 24-hour cytotoxicity study evaluates only acute effects; thus, extended exposure studies (48–72 h) are needed to determine chronic cytotoxicity. Further research is also required to test the nanoparticles against polymicrobial biofilms commonly present in endodontic infections. In vivo studies are necessary to validate the safety and efficacy of biosurfactant-coated CuNPs for clinical applications.
Conclusions
This study successfully synthesized highly pure and stable biosurfactant-coated CuNPs using biosurfactants derived from A. xylosoxidans and P. aeruginosa as biostabilizers. Coating with biosurfactants significantly enhanced the stability, morphology and antimicrobial efficacy of the nanoparticles. Characterization analyses confirmed their spherical morphology, with particle sizes ranging from 90 nm to 100 nm. Zeta potential measurements indicated moderate stability and biocompatibility.
CuO+P5C and CuO+P11 demonstrated potent antimicrobial activity against key endodontic pathogens (E. coli, S. aureus, S. oralis, and Enterococcus spp.), achieving a MIC of 125 µg/mL. The cytotoxicity evaluation on fibroblast cell lines (L929) revealed minimal toxicity, with nearly 78–80% cell viability maintained at concentrations up to 100 µg/mL, underscoring their excellent biocompatibility.
The eco-friendly and cost-effective use of biosurfactants as stabilizers positions these nanoparticles as a promising alternative to chemical-based antimicrobials for addressing persistent endodontic infections. Future in vivo studies and extended cytotoxicity evaluations are essential to validate their clinical efficacy and ensure their safe application in medical and dental fields.
Ethics approval and consent to participate
Not applicable.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
Consent for publication
Not applicable.
Use of AI and AI-assisted technologies
Not applicable.