Abstract
Background. Fragile X-related protein 1 (FXR1), an RNA-binding protein (RBP) implicated in various cancers, has emerged as a potential contributor to the progression of head and neck squamous cell carcinoma (HNSCC).
Objectives. The aim of the study was to comprehensively analyze FXR1 genetic alterations, mRNA expression, protein levels, and immune infiltration in HNSCC, in order to elucidate its molecular role and therapeutic potential.
Material and methods. Data from The Cancer Genome Atlas (TCGA)–HNSCC cohort was analyzed. Copy number variation (CNV) was assessed using cBioPortal, FXR1 expression profiling was performed using the University of ALabama at Birmingham CANcer data analysis Portal (UALCAN), and immunohistochemistry (IHC) examination was conducted using the Human Protein Atlas (HPA). The TIMER 2.0 and TISIDB databases were employed for immune infiltration analysis.
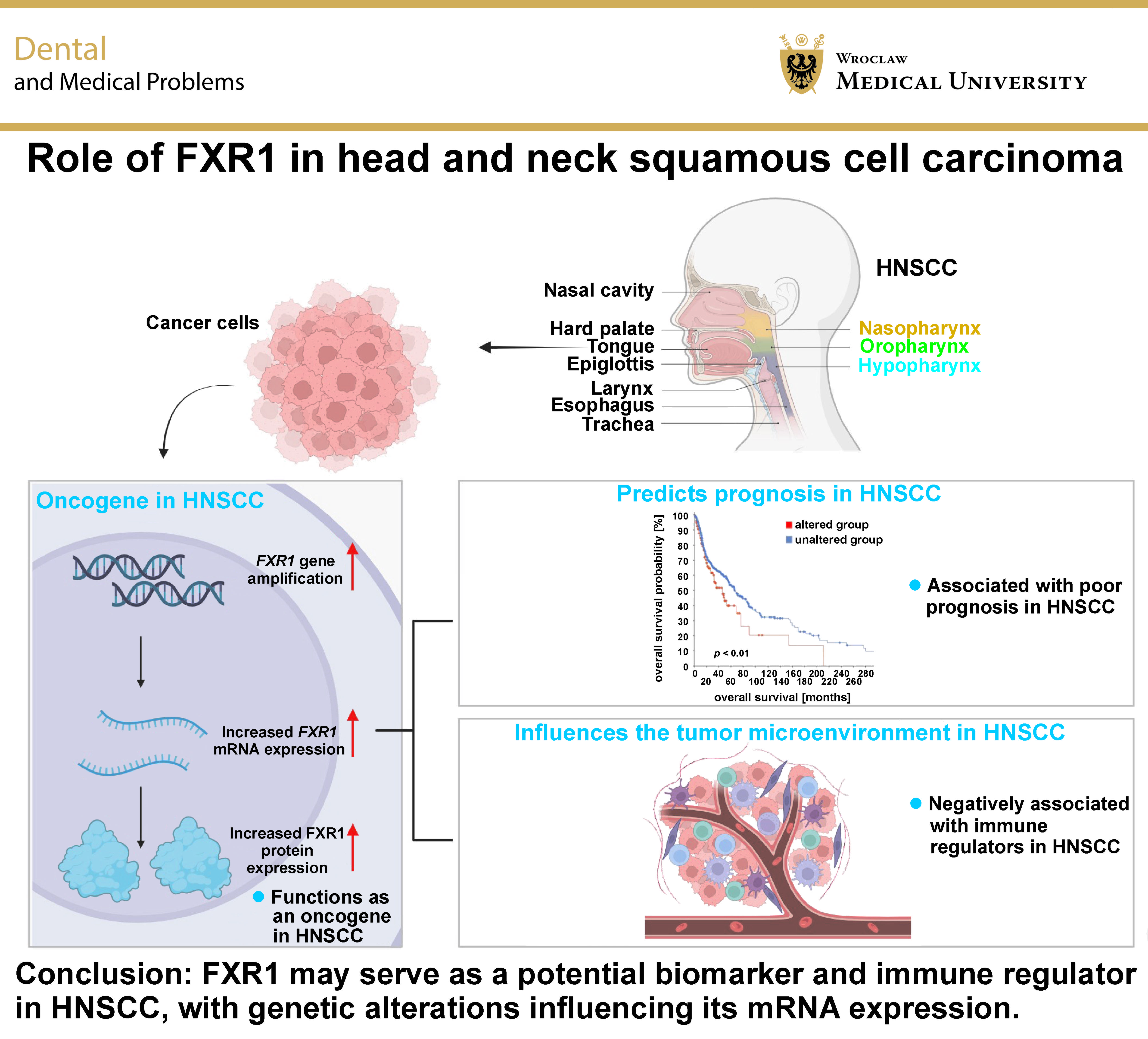
Results. Genetic alterations in FXR1, including DNA amplification, were observed in 22% of HNSCC patients and were significantly correlated with elevated mRNA expression. FXR1 mRNA and protein levels were upregulated in HNSCC tumors, and this upregulation was associated with poor prognosis. The expression of FXR1 was also correlated with clinical features, including disease stage, grade, metastasis, and human papillomavirus (HPV) status. Immune infiltration analysis revealed a negative correlation between FXR1 mRNA expression and CD8+ T cells, suggesting a potential role in immune modulation.
Conclusions. FXR1 may serve as a potential biomarker and immune regulator in HNSCC, with genetic alterations influencing its mRNA expression. This study sheds light on the multifaceted role of FXR1 in the progression of HNSCC, and suggests potential avenues for targeted therapeutic strategies. Further research, including functional experiments and large-scale cohort analyses, is warranted to validate and translate these findings into clinical applications.
Keywords: TCGA, prognosis, copy number variation, head and neck carcinoma, FXR1
Introduction
Head and neck squamous cell carcinoma (HNSCC) constitutes a diverse group of tumors, predominantly squamous cell carcinomas (SCCs), accounting for approx. 90% of diagnoses and ranking as the 6th most prevalent cancer worldwide.1, 2 Oropharyngeal cancers are primarily driven by human papillomavirus (HPV), while alcohol consumption and tobacco use are risk factors for cancers of the oral cavity, hypopharynx and larynx. Notably, HPV-positive HNSCC generally exhibits a more favorable prognosis.3 The development of HNSCC involves intricate genetic and epigenetic alterations that contribute to hallmark features of cancer, including uncontrolled proliferation and metastasis.4, 5, 6, 7, 8
The discovery of human fragile X-related protein 1 (FXR1) (3q28) resulted from fragile X messenger ribonucleoprotein 1 (FMR1) sequencing, revealing an autosomal gene with 60% amino acid sequence identity to other family members.9 FXR1 shares a common structure but differs in its C-termini, implying distinct roles. The protein contains several domains, including a Tandem agent-like domain, a non-classical nuclear localization signal (NLS), 3 K homology (KH) domains, a nuclear export signal (NES), and an arginine–glycine–glycine (RGG) box, classifying FXR1 as an RNA-binding protein (RBP). Exportin-1 (XPO-1) shuttles FXR1 to the cytoplasm, where RGG motifs facilitate binding to DNA or RNA structures. FXR1 is ubiquitously expressed and undergoes various post-translational modifications, including SUMOylation, ubiquitylation, acetylation, methylation, and phosphorylation at S420.10, 11
FXR1 regulates microRNA (miRNA) processing, stabilizes specific miRNAs, and influences their subcellular localization.12 It interacts with nucleoporins during mitosis, affecting protein export. Additionally, FXR1 stabilizes mRNA through adenylate–uridylate (AU)-rich elements (AREs) in the 3’-untranslated region (3’-UTR), thereby influencing mRNA translation and stability. It also modulates mRNA translation through interactions with miRNAs, argonaute-2 (AGO2) and the RNA interference (RNAi) pathway. Furthermore, FXR1 has an influence on the cell cycle, synaptic protein function and muscle development. It plays a crucial role in tumor necrosis factor alpha (TNF-α)–ARE-mediated translational activation, impacting cell growth. Overall, these multifaceted functions highlight the significance of FXR1 in post-transcriptional gene regulation and underscore its potential diagnostic and therapeutic implications.10, 13
Recent studies have highlighted the involvement of FXR1 in cancer.14, 15 Elevated expression of FXR1 has been observed in various malignancies, including bladder cancer, colorectal cancer, non-small cell lung cancer, and hepatocellular carcinoma (HCC).14, 15, 16, 17, 18 In these contexts, FXR1 acts as an oncogene, promoting tumor growth and invasion through mechanisms such as mRNA stabilization, enhanced protein translation and inactivation of tumor suppressor genes.19, 20, 21
Despite its pro-tumorigenic attributes, the RNA-binding mechanisms of FXR1 offer promising therapeutic avenues. Targeting these interactions using small molecules or RNAi-based strategies may disrupt its oncogenic effects. In HNSCC, genetic alterations, particularly copy number variation (CNV), play a crucial role in activating oncogenes or inactivating tumor suppressor genes.22, 23, 24 Recent studies have also demonstrated the overexpression of FXR1 in HNSCC tissues, correlating it with tumor growth and invasion.25 This study, utilizing data from The Cancer Genome Atlas (TCGA)–HNSCC cohort, aimed to analyze FXR1 genetic variations, including the expression of DNA, mRNA, protein, as well as to unravel its molecular role and therapeutic potential in HNSCC. FXR1 is significantly associated with HNSCC, thus providing a link between the prognosis and tumor infiltration in HNSCC.
Material and methods
Copy number variation analysis
The cBioPortal (https://www.cbioportal.org), a user-friendly and interactive online resource, facilitates the visualization, analysis and download of extensive cancer genomics datasets.26 In this investigation, the cBioPortal was used to examine alterations in the FXR1 gene among HNSCC patients. Comprehensive clinical data for 528 HNSCC patients is presented in Table 1. The prognostic significance of FXR1 genetic alterations in patients with HNSCC was assessed using the Kaplan–Meier plotter analysis.
FXR1 expression analysis
The University of ALabama at Birmingham CANcer data analysis Portal (UALCAN) (http://ualcan.path.uab.edu)27 was used to analyze expression profiles in HNSCC. The UALCAN integrates datasets from TCGA and the Clinical Proteomic Tumor Analysis Consortium (CPTAC). The present study also involved a comprehensive analysis of the relationship between FXR1 expression and clinicopathological features of HNSCC, such as disease stage, histological grade, lymph node involvement, and HPV status.
Immunohistochemistry analysis
The Human Protein Atlas (HPA) (http://www.proteinatlas.org)28 is a database that provides information on human protein-coding genes, including RNA and protein expression levels as well as location. Immunohistochemistry (IHC) images illustrating the expression of FXR1 protein in clinical specimens from HNSCC patients and normal tissues were obtained from the HPA database. To analyze the expression levels of FXR1, staining intensity was classified as low, moderate or high. According to HPA data, Anti-FXR1 rabbit monoclonal antibodies (HPA018246 and HPA055475; Sigma-Aldrich, St. Louis, USA) were utilized for IHC, with dilution ratios of 1:150 and 1:50, respectively.
Immune infiltration analysis
Given the pivotal role of immune genes in the pathogenesis of HNSCC, we analyzed the relationship between our target gene, FXR1, and key immune genes involved in HNSCC pathogenesis. Tumor infiltration was assessed using TIMER 2.0 (http://TIMER.cistrome.org)29 and the TISIDB (http://cis.hku.hk/TISIDB)30 database. The TIMER server was employed to systematically evaluate immune infiltration across various malignancies, employing the TIMER method to quantify CD4+ T cells, CD8+ T cells, B cells, neutrophils, macrophages, and dendritic cells. The gene module was applied to examine FXR1 expression in different cancers and the extent of immune cell infiltration in HNSCC. Additional analyses included the exploration of immunomodulators, chemokines and their receptors. Specifically, the top 3 negatively correlated immunomodulatory genes, encompassing major histocompatibility complex (MHC) molecules, immunostimulators and immune inhibitors, were identified in relation to FXR1 mRNA expression in HNSCC patients.
Statistical analysis
Data was analyzed using various statistical methods depending on the specific research question. To compare FXR1 expression between the groups, Student’s t-tests were performed to calculate p-values and determine fold changes, along with gene rank information. The Kaplan–Meier analysis was used to assess the relationship between variables and survival outcomes, with the log-rank test applied for comparisons between survival curves. To comprehensively evaluate the impact of FXR1 expression and other clinicopathological factors on survival, a multivariate Cox proportional hazards model was employed. Correlations between gene expression levels were assessed using Spearman’s correlation analysis in the TIMER 2.0 databases. The TISIDB database provided both Spearman’s and Pearson’s correlation analyses to evaluate the strength of association between variables. A p-value <0.05 was considered statistically significant unless stated otherwise.
Results
Association between genetic alterations in FXR1 and increased mRNA expression in HNSCC
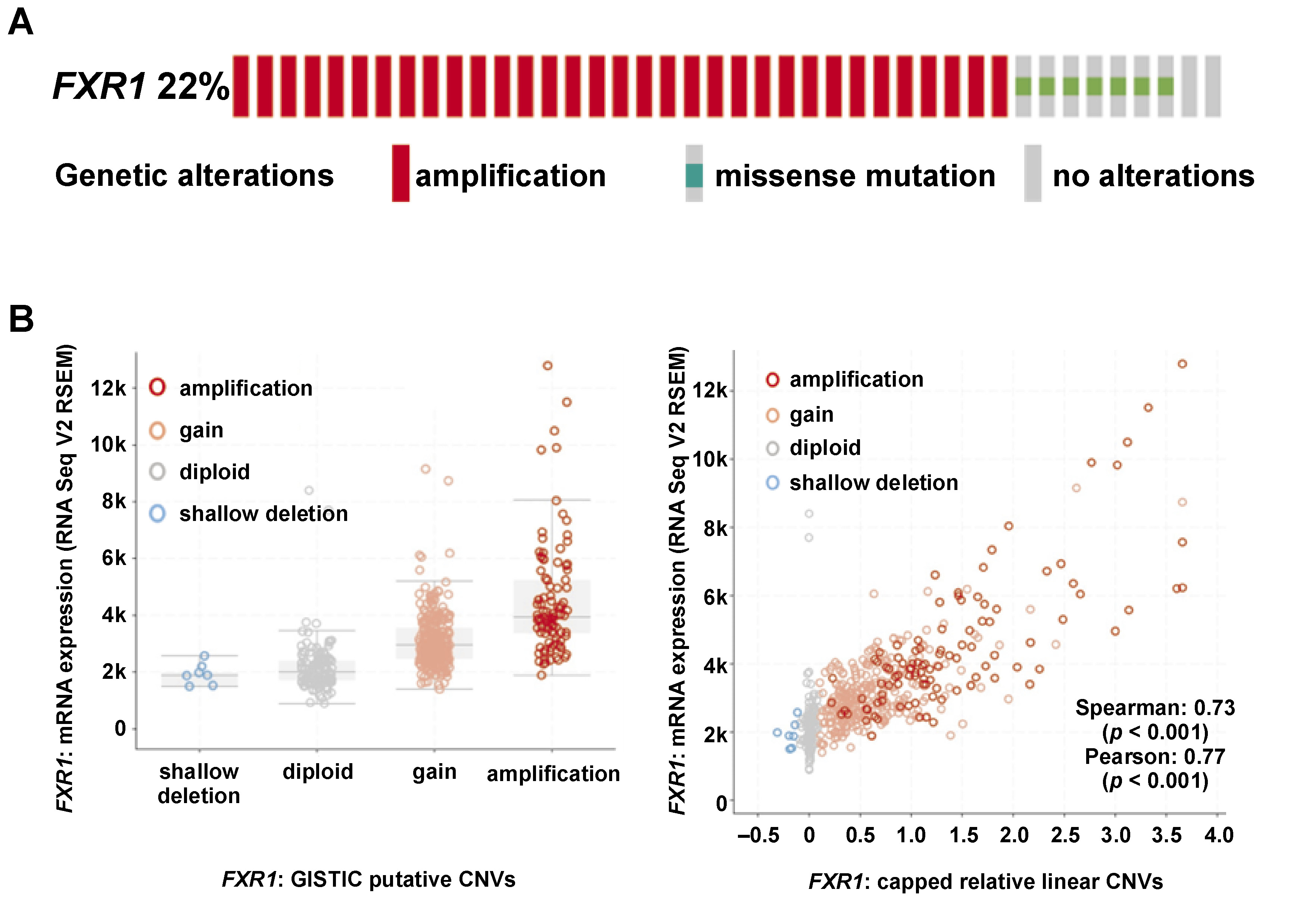
Genetic alterations in FXR1 were observed in 22% of HNSCC patients among the 528 individuals analyzed. These alterations were primarily characterized by FXR1 gene amplification, while some cases exhibited missense mutations (Figure 1A). Notably, FXR1 amplification was significantly correlated with elevated FXR1 mRNA expression in HNSCC patients (Figure 1B) (p < 0.05).
Upregulation of FXR1 mRNA and protein levels in HNSCC
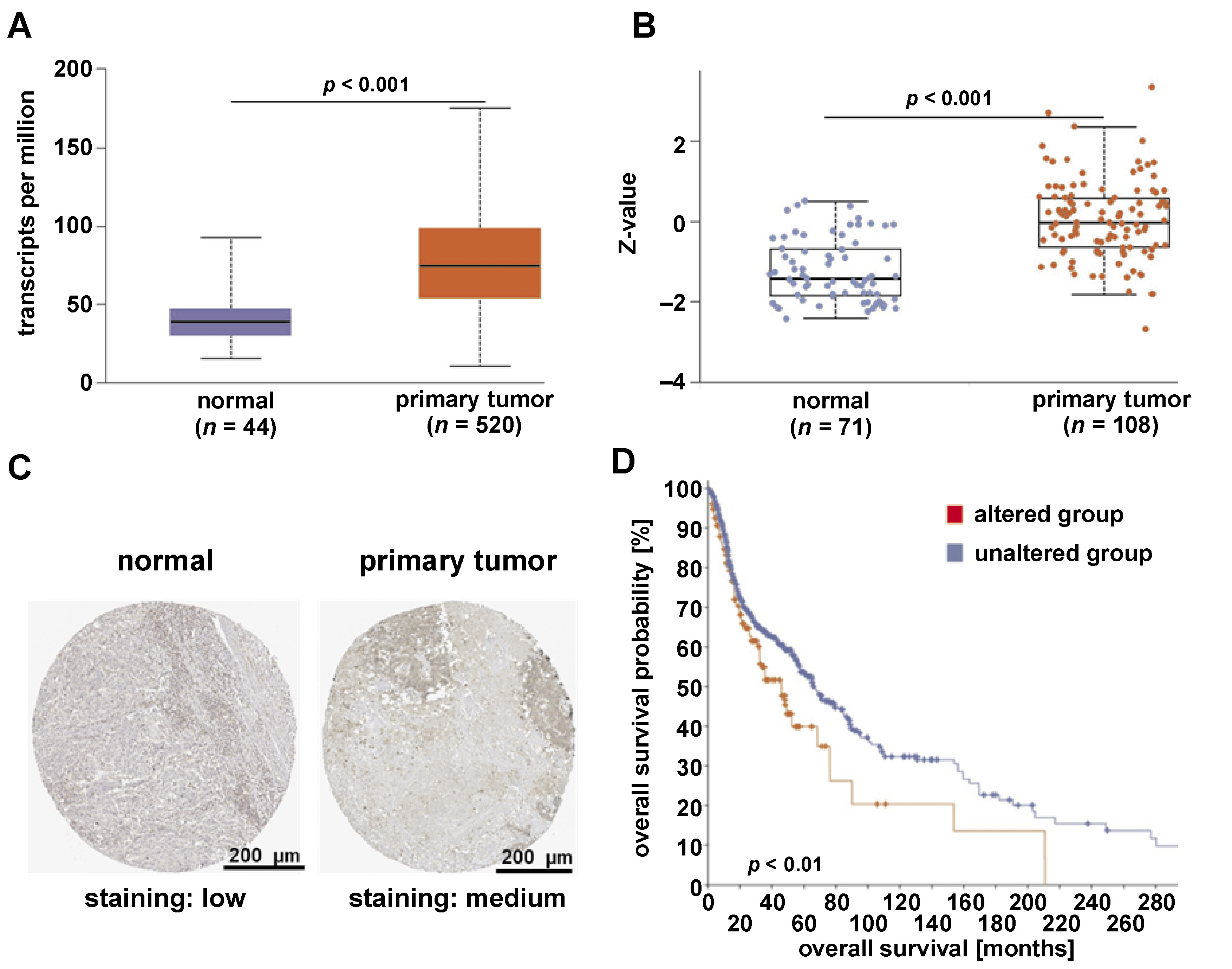
The mRNA expression level of FXR1 was significantly upregulated in patients with primary HNSCC tumors (n = 520) compared to normal tissues (n = 44) in the TCGA–HNSCC dataset (Figure 2A) (p < 0.001). Similarly, FXR1 protein levels were significantly increased in primary HNSCC tumor tissues (n = 108) compared to normal tissues (n = 71) in the CPTAC–HNSCC dataset (Figure 2B) (p < 0.001). Immunohistochemical analysis further confirmed that FXR1 protein was markedly higher in HNSCC tumor tissues than in normal tissues (Figure 2C). Moreover, the Kaplan–Meier curve displayed the relationship between FXR1 genetic alterations and HNSCC patient survival. In particular, patients with altered FXR1 exhibited poorer overall survival, suggesting its potential prognostic significance (Figure 2D) (p < 0.05).
Association of dysregulated FXR1 mRNA with stage, grade, metastasis status, and HPV status in HNSCC
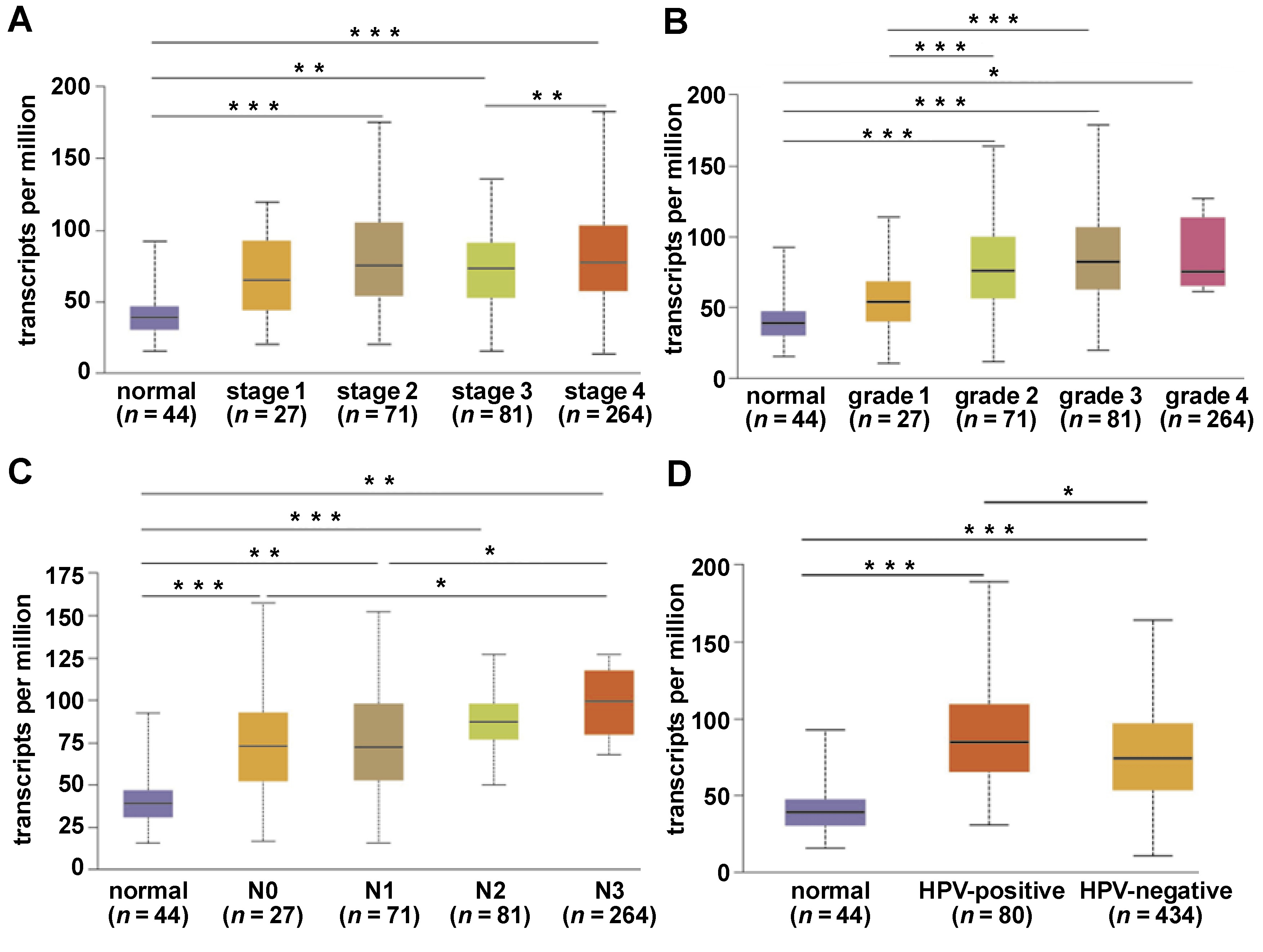
FXR1 mRNA expression was significantly altered across different tumor stages, grades and metastasis statuses compared to normal tissues (Figure 3A–C) (p < 0.05). In terms of tumor stage, significant differences were noted between normal tissues and stages 2, 3 and 4, as well as between stages 3 and 4 (Figure 3A) (p < 0.05). Regarding HNSCC grade, substantial alterations were observed between normal tissues and grades 2, 3 and 4, as well as between grade 1 and grades 2 and 3 (Figure 3B) (p < 0.05). In terms of metastasis status, FXR1 expression in normal tissues differed significantly from N0–N3 groups. Additionally, significant differences were observed between N0 and N3, and between N1 and N3 (Figure 3C) (p < 0.05). Furthermore, FXR1 expression was significantly higher in HPV-positive HNSCC patients compared to HPV-negative patients and normal tissues (Figure 3D) (p < 0.05).
Impact of FXR1 mRNA expression on tumor immune infiltration in HNSCC
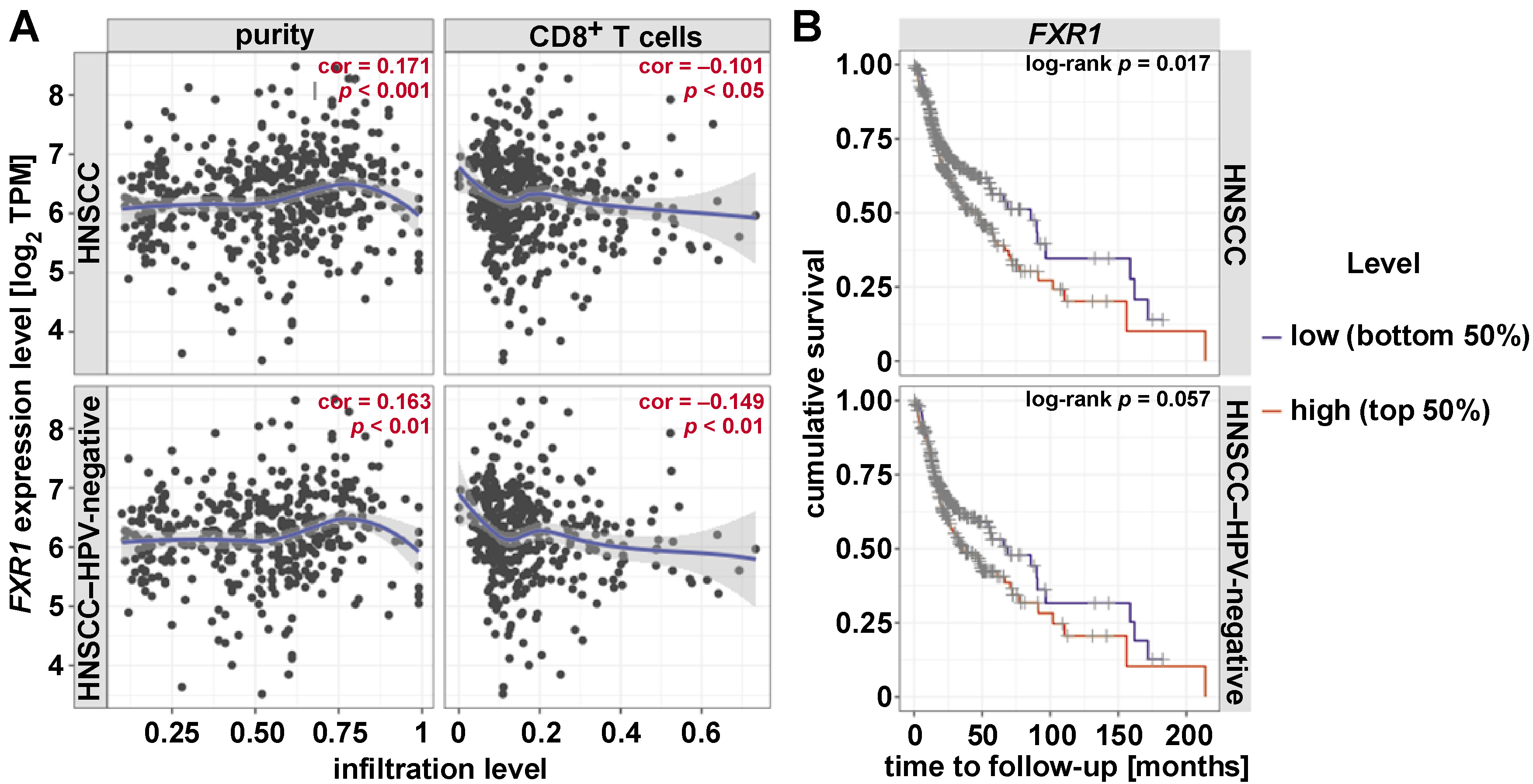
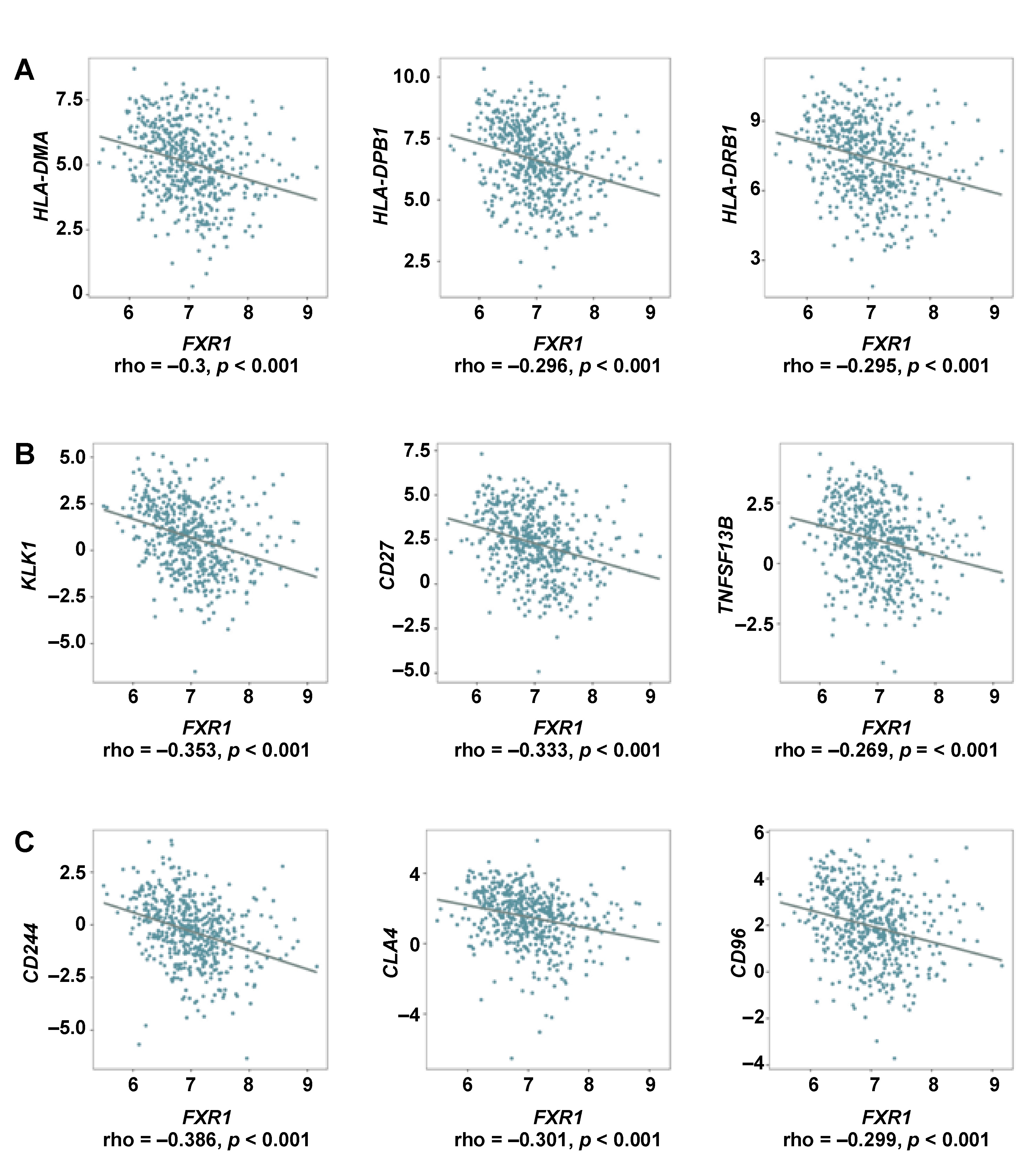
The mRNA expression of FXR1 exhibited a significant negative correlation with CD8+ T cells (Figure 4A). High FXR1 expression was associated with poorer prognosis in HNSCC patients, including those with HPV-negative tumors (Figure 4A,B). The mRNA expression of FXR1 and genes involved in MHC molecules, immunostimulators and immunoinhibitors, were analyzed in HNSCC patients, as depicted in Supplementary Figure 1 (available on request from the corresponding author). These groups function as immunomodulators in humans. The top 3 negatively correlated genes in each category were identified, with MHC molecules (HLA-DMA, HLA-DPB1, HLA-DRB1), immunostimulators (KLRK1, CD27, TNFSF13B) and immune inhibitors (CD244, CTLA4, CD96) showing significant negative correlations with FXR1 expression in HNSCC patients (Figure 5A–C) (p < 0.001).
Discussion
Among RNA-binding proteins, FXR1 is a well-characterized oncogene involved in multiple cancers. The overexpression of FXR1 has been reported in several malignancies, including glioma, lung carcinoma, HCC, breast cancer, head and neck carcinoma, ovarian cancer, and colorectal carcinoma.14, 15, 17, 18, 20, 25
Copy number variation has been associated with changes in gene expression; yet, the specific cellular associations between CNV and gene expression alterations remain unknown.31 The present study investigated genetic alterations and their functional implications, as well as the association between FXR1 mRNA expression and HNSCC. Genetic alterations in FXR1 were observed in 22% of HNSCC patients and were primarily characterized by gene amplification, with a smaller proportion of cases exhibiting missense mutations. Notably, FXR1 amplification was significantly correlated with elevated FXR1 mRNA expression, indicating a potential regulatory mechanism. This observation confirms previous findings demonstrating the influence of gene copy number on expression levels. Additionally, earlier studies have reported enrichment of both amplified and downregulated genes within the ubiquitin–proteasome system, suggesting potential consequences for protein degradation pathways in HNSCC.31 Further research by Baltaci et al.32 and Carron et al.33 demonstrated that CNVs, particularly gene amplification, contribute to HNSCC carcinogenesis through the cooperation of oncogenes and inherited CNV, influencing the occurrence and prognosis of oropharyngeal cancer, respectively. Beyond oncology, genetic alterations in FXR1 have also been implicated in congenital multiminicore myopathy and Fragile X syndrome.34, 35
FXR1 emerges as a potential biomarker and immune regulator in HNSCC. Both mRNA and protein levels were elevated in tumor tissues compared to healthy tissues, suggesting a role in disease progression. This upregulation is associated with patient survival and correlates with diverse clinical features, including HPV status. Previous studies have examined the role of FXR1 in cancer. Xiao et al. demonstrated the importance of FXR1 across 25 cancer types, including breast and cervical cancers, showcasing its diagnostic potential and association with prognosis.36 The enrichment analysis revealed the influence of FXR1 on tumor development and progression through multiple pathways such as cell cycle control, mitochondrial metabolism, immune response, and skin barrier formation.36 Notably, RBPs have emerged as key regulators of these processes, influencing RNA splicing, transport, translation, and localization.37 Further studies have demonstrated the diverse roles of FXR1, ranging from influencing RNA metabolism in glioblastoma to impacting muscle development and adenovirus interactions.20, 38, 39 These findings reveal the broad involvement of FXR1 in various cancer types and disease-related processes.
While existing research explores the role of FXR1 in cancer, several key mechanisms remain shrouded. In ovarian cancer, FXR1 has been shown to regulate c-MYC translation and stabilization, promoting its oncogenic activity.40 Moreover, cancer cells lacking both p53 and FXR2 (“double knockout”) exhibit increased sensitivity to FXR1 inhibition. This vulnerability depends on the FXR1 tandem Tudor domain and its interaction with STAT1/3 proteins. FXR1 binds to gene promoters, including ARID1A and CASC4, and recruits STAT1/3 to promote transcription. The inhibition of FXR1 or JAK (activating STAT1/3) suppresses FXR1 target genes and reduces cancer cell growth.41 These findings reveal a novel synergy between FXR1 and STAT1/3 in regulating cell proliferation, opening therapeutic avenues for double-knockout cancers. Furthermore, in HCC, FXR1 has been implicated in the intricate transforming growth factor beta (TGF-β)/SMAD signaling pathway. Elevated FXR1 expression correlates with dismal prognosis and promotes cancer cell motility, invasion and proliferation through the modulation of the SMAD2/3 axis.18 Moreover, Majumder et al. highlighted the ability of FXR1 to bypass cellular senescence in oral cancer, revealing its contribution to tumor cell survival and growth.42 These outcomes suggest that FXR1 presents additional therapeutic opportunities across diverse cancer landscapes.
A growing number of studies have shown that the immune microenvironment plays a vital role in the development and progression of HNSCC.43 Previous studies have shown that FXR1 expression is significantly correlated with the expression of immunomodulators across various cancers.36 In this study, FXR1 expression was negatively correlated with CD8+ T cells, implying a role in the modulation of the immune microenvironment. Further analyses indicated that FXR1 interacts with key immune-related genes, supporting its potential as an immunotherapeutic target. These findings warrant further investigation into the multifaceted role of FXR1 in HNSCC progression and immune evasion.
The current study, based on the TCGA–HNSCC dataset and various bioinformatics tools, provides valuable insights into the potential of FXR1 as a biomarker and prognostic indicator in HNSCC. However, further research is required to fully grasp its potential. Large-scale cohort analysis of genetic variants and expression patterns across diverse experimental methods could significantly enhance the accuracy and robustness of these findings. Functional experiments and in vivo studies are crucial for validating the therapeutic potential of FXR1 in HNSCC. Our initial data has shed light on how genetic variants focusing on CNV influence the pathogenesis of HNSCC by altering FXR1 expression and regulation. Building upon this foundation, further research can translate these findings into tangible therapeutic strategies.
Conclusions
This study demonstrates a significant association between FXR1 genetic alterations and aberrant expression in HNSCC. FXR1 gene amplification contributes to elevated FXR1 mRNA and protein levels in HNSCC patients. This upregulation is associated with poor prognosis and correlates with various clinical features, including disease stage, grade, lymph node metastasis status, and HPV status. Additionally, our findings suggest that FXR1 may play a role in modulating the immune microenvironment, emphasizing its multifaceted involvement in HNSCC progression and immune evasion. Taken together, FXR1 emerges as a promising biomarker and therapeutic target, warranting further exploration of its molecular mechanisms and therapeutic potential in HNSCC.
Ethics approval and consent to participate
Not applicable.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
Consent for publication
Not applicable.
Use of AI and AI-assisted technologies
ChatGPT (OpenAI, San Francisco, USA) was used to improve readability. The content was then subjected to a thorough review and editing process to ensure the highest standards of quality. The authors accept full responsibility for the publication.